Mr. Bean App

In plant improvement experimental designs are used to decompose the total phenotypic variance observed in field experiments into at least two components: A genetic and a non-genetic component that is attributable to the spatial variation or environment. Recently, new methodologies for the modelling of spatial trends have been published using the arrangement of the experimental units in the field. These methodologies have shown an improvement in the prediction of the genetic potential of evaluated genotypes.
However, the use of these tools may be limited because of the cost to access a licensed product and/or the requirement to be familiar with the language and environment that was used for their implementation. This, in turn limits the data analysis efficiency for decision making. These limitations led to the development of Mr.Bean, an easy-to-access, practical and user-friendly tool that integrates the spatial modelling capabilities of SpATS, the graphical versatility of plotly and the interactive and simple construction approach offered by Shiny for the development of Web applications.
Features are built-in separate modules, and users can start to discover the advantages of this tool as soon as they download the package!
How to Install MrBean: An app for experimental trial analysis using Linear Mixed Models
In what context is this tool useful?
Mr. Bean app is useful in a context where is need it a convenient and accurate way to analyze agronomic data, visualize field patterns and select genotypes all in one place using the latest statistical and computational tools.
This tool incorporates descriptive analyses, measures of dispersion and centralization, graphical visualization for comparing multiple variables, the adjustment of mixed models with or without spatial components and the identification of outlier data. All these capabilities are aimed at plant breeders and in general people working with agricultural field data to make precise decisions more quickly.
Schematic representation of Mr.Bean’s modules
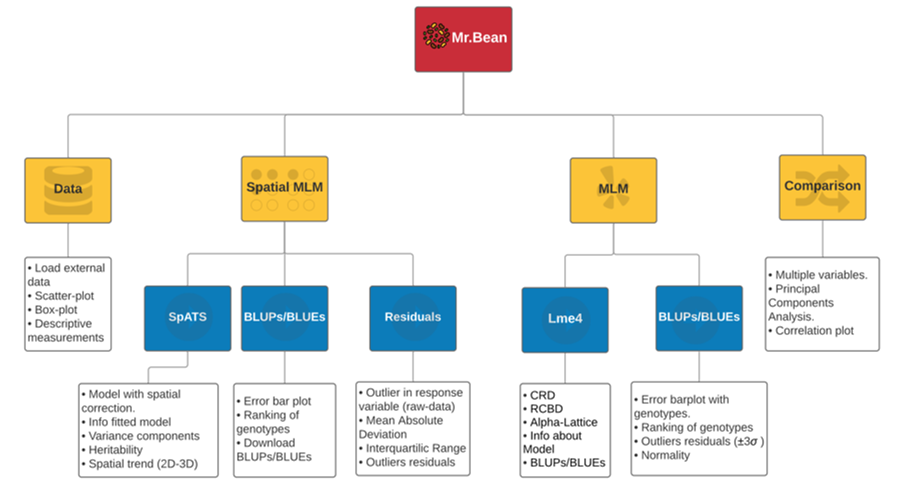
Expected Results
Mr. Bean app results are BLUPs/BLUEs predictions and heritabilities for single-environmental experiments or multiple-environmental trial (MET) analysis. In addition, Mr. Bean also provides a module for exploring results from METs using several graphical and multivariate techniques.
With Mr. Bean, users can obtain adjusted means for gene numbers that can be used in determining Genetic Gain, recycling, and selection decisions. This app also makes easier to combine phenotypic and genotypic data with a GBLUP module available that allows that users get:
- Marker Effects.
- Variance Components.
- Genomic Predictions.
- Marker-based heritabilty (genomic heritability).
- GBLUPs.
- Genotypic Correlations.
Contact people
Johan Steven Aparicio – [email protected]

